Research Activities
- Publication List [Research Activity Database]
Recent Studies
1. Direct numerical simulations (DNS) for colloidal dispersions
We have developed a unique mesoscale method for simulating colloidal dispersions. Our program has been released as a colloid simulator KAPSEL, which enables us to perform successful DNS simulations for neutral and charged colloidal dispersions. We have applied this method to analyze the dynamics of self-propelled particles for a schematic model of micro-organisms.
2. Development of multiscale simulation method to predict polymeric liquid flows and its applications to industrial polymer processing
To manufacture more sophisticated polymeric products, it is needed to predict and control flow behaviors of polymeric liquids in industrial processes. In general, it is difficult to do this for polymeric chains with a molecular weight distribution and with various types of polymer architecture, because a microscopic state of polymer chains such as an orientation and entanglements of constituent polymer chains does strongly influence on the macroscopic flow behavior. To deal with macro-scopic flow behaviors and micro-scopic state of polymer chains simultaneously, it has been desired strongly to establish a multiscale simulation (MSS) method that enables to bridge between the two different spatial scales. For this purpose, we are addressing to develop the MSS method and to apply it to various industrial polymer processing. (See Fig.1: application of the MSS method to a polymer melt spinning process).
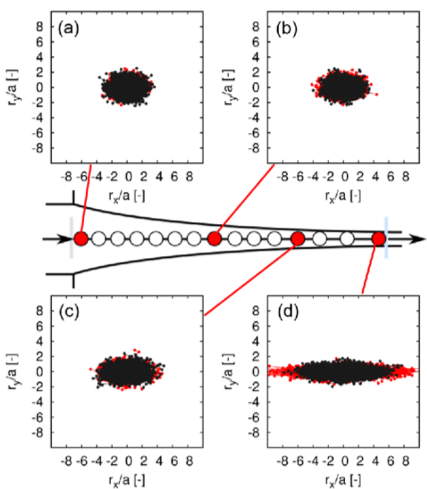
Fig. 1. Configuration of polymer chains at four points (a)-(d) on a thread of a polymer melt spinning process. Red: longer polymer chain, Black: shorter polymer chain.
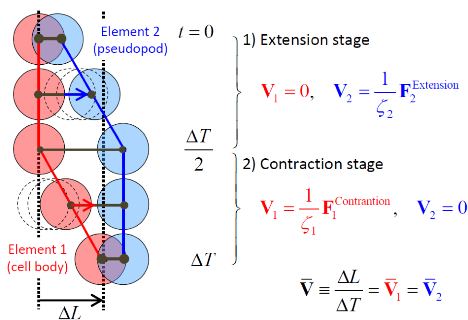
3. Physical modeling of biological tissues
Inspired by recent studies on the model of crawling and dividing cells [Coburn et al., Phys. Biol. 10, 046002 (2013); Basan et al., PNAS 110, 2452 (2013).], we developed a unique particle-based minimal model for crawling and dividing cells on substrate (see Fig. 2). It mimics a real mechanics of migrating/dividing cells with the mechanisms of the contact inhibition of locomotion (CIL) and the contact inhibition (CI) of cell division in a straightforward way. The present model has been applied to simulate the dynamics of growing colony composed of active cells. Some basic properties seen in real growing colonies have been successfully reproduced. The true mechanism behind such complex biological systems will be discussed in physical context.